Expression Profile of PmiR-31, Novel npcRNA Of Proteus Mirabilis Under Different Growth Phases and Stress Conditions
DOI:
https://doi.org/10.22452/mjs.vol44sp1.4Keywords:
Proteus mirabilis, non-protein coding RNA, PmiR-31, flagella biosynthesisAbstract
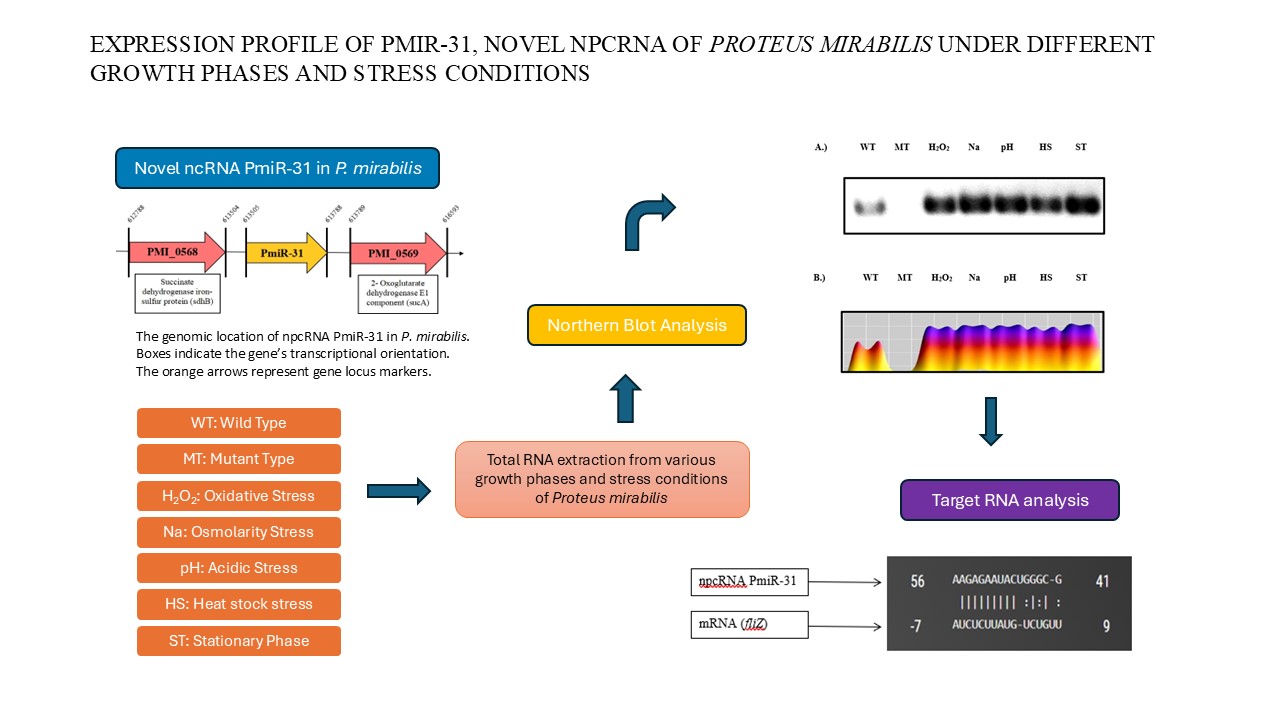
Proteus mirabilis (P. mirabilis), known for its swarming motility, is a facultatively anaerobic, rod-shaped, Gram-negative bacterium. It forms extended swarmer cells as well as moves in a bull’s-eye formation. Proteus mirabilis significantly causes catheter-associated urinary tract infections through virulence components including flagella, fimbriae, hemolysin, urease, and proteases. Flagella-related motility allows P. mirabilis to infect the urinary system. The flagellum, composed of about 20 proteins, has a body that penetrates the cell wall, a curved hook, and a filament extending several micrometers. Previously, we knocked out PmiR-137, anticipated to modulate flhDC, which involve in the central transcriptional activator for flagellum formation and performed a differential gene expression study. We discovered npcRNA PmiR-31, expected to influence fliZ production via the TargetRNA web tool. We analyzed PmiR-31 expression under various stress and growth phases via northern blot. The PmiR-31 was highly expressed in stress conditions but not in the mutant strain. Since FliZ regulates flagella assembly, npcRNA PmiR-31 may repress fliZ translation, preventing flagella synthesis and promoting biofilm formation to evade the immune system.
References
Akahoshi, D. T., & Bevins, C. L. (2022). Flagella at the Host-Microbe Interface: Key Functions Intersect With Redundant Responses. Frontiers in Immunology, 13, 828758. https://doi.org/10.3389/fimmu.2022.828758
Avelino-Flores, F., Soria-Bustos, J., Saldaña-Ahuactzi, Z., Martínez-Laguna, Y., Yañez-Santos, J. A., Cedillo-Ramírez, M. L., & Girón, J. A. (2022). The Transcription of Flagella of Enteropathogenic Escherichia coli O127:H6 Is Activated in Response to Environmental and Nutritional Signals. Microorganisms, 10(4), Article 4. https://doi.org/10.3390/microorganisms10040792
Bientz, V., Lanois, A., Ginibre, N., Pagès, S., Ogier, J.-C., George, S., Rialle, S., & Brillard, J. (2024). OxyR is required for oxidative stress resistance of the entomopathogenic bacterium Xenorhabdus nematophila and has a minor role during the bacterial interaction with its hosts. Microbiology, 170(7), 001481. https://doi.org/10.1099/mic.0.001481
Chakkour, M., Hammoud, Z., Farhat, S., El Roz, A., Ezzeddine, Z., & Ghssein, G. (2024). Overview of Proteus mirabilis pathogenicity and virulence. Insights into the role of metals. Frontiers in Microbiology, 15, 1383618. https://doi.org/10.3389/fmicb.2024.1383618
Chinni, S. V., Raabe, C. A., Zakaria, R., Randau, G., Hoe, C. H., Zemann, A., Brosius, J., Tang, T.-H., & Rozhdestvensky, T. S. (2010). Experimental identification and characterization of 97 novel npcRNA candidates in Salmonella enterica serovar Typhi. Nucleic Acids Research, 38(17), 5893–5908. https://doi.org/10.1093/nar/gkq281
Das, C., Mokashi, C., Mande, S. S., & Saini, S. (2018). Dynamics and Control of Flagella Assembly in Salmonella typhimurium. Frontiers in Cellular and Infection Microbiology, 8. https://doi.org/10.3389/fcimb.2018.00036
Eran, Z., Akçelik, M., Yazıcı, B. C., Özcengiz, G., & Akçelik, N. (2020). Regulation of biofilm formation by marT in Salmonella Typhimurium. Molecular Biology Reports, 47(7), 5041–5050. https://doi.org/10.1007/s11033-020-05573-6
Filipiak, A., Chrapek, M., Literacka, E., Wawszczak, M., Głuszek, S., Majchrzak, M., Wróbel, G., Gniadkowski, M., & Adamus-Białek, W. (2020). Correlation of pathogenic factors with antimicrobial resistance of clinical Proteus mirabilis strains (p. 2020.02.24.962514). bioRxiv. https://doi.org/10.1101/2020.02.24.962514
Gmiter, D., & Kaca, W. (2022). Into the understanding the multicellular lifestyle of Proteus mirabilis on solid surfaces. Frontiers in Cellular and Infection Microbiology, 12. https://doi.org/10.3389/fcimb.2022.864305
Hengge, R. (2020). Linking bacterial growth, survival, and multicellularity – small signaling molecules as triggers and drivers. Current Opinion in Microbiology, 55, 57–66. https://doi.org/10.1016/j.mib.2020.02.007
Hengge, R., Pruteanu, M., Stülke, J., Tschowri, N., & Turgay, K. (2023). Recent advances and perspectives in nucleotide second messenger signaling in bacteria. MicroLife, 4, uqad015. https://doi.org/10.1093/femsml/uqad015
Jamil, R. T., Foris, L. A., & Snowden, J. (2025). Proteus mirabilis Infections. In StatPearls. StatPearls Publishing. http://www.ncbi.nlm.nih.gov/books/NBK442017/
Kędzierska-Mieszkowska, S. (2023). Sigma factors of RNA polymerase in the pathogenic spirochaete Leptospira interrogans, the causative agent of leptospirosis. The FASEB Journal, 37(10), e23163. https://doi.org/10.1096/fj.202300252RRR
Kim, S. W., Li, Z., Moore, P. S., Monaghan, A. P., Chang, Y., Nichols, M., & John, B. (2010). A sensitive non-radioactive northern blot method to detect small RNAs. Nucleic Acids Research, 38(7), e98. https://doi.org/10.1093/nar/gkp1235
Kishanraj, S., Sumitha, S., Tang, T.-H., Citartan, M., & Chinni, S. V. (2021a). Comparative genomic identification and characterization of npcRNA homologs in Proteus vulgaris. Journal of Biosciences, 46, 108.
Kishanraj, S., Sumitha, S., Tang, T.-H., Citartan, M., & Chinni, S. V. (2021b). Comparative genomic identification and characterization of npcRNA homologs in Proteus vulgaris. Journal of Biosciences, 46, 108.
Kurniyati, K., Chang, Y., Guo, W., Liu, J., Malkowski, M. G., & Li, C. (2023). Anti-σ28 Factor FlgM Regulates Flagellin Gene Expression and Flagellar Polarity of Treponema denticola. Journal of Bacteriology, 205(2), e00463-22. https://doi.org/10.1128/jb.00463-22
Ponath, F., Hör, J., & Vogel, J. (2022). An overview of gene regulation in bacteria by small RNAs derived from mRNA 3’ ends. FEMS Microbiology Reviews, 46(5), fuac017. https://doi.org/10.1093/femsre/fuac017
Raabe, C. A., Sanchez, C. P., Randau, G., Robeck, T., Skryabin, B. V., Chinni, S. V., Kube, M., Reinhardt, R., Ng, G. H., Manickam, R., Kuryshev, V. Y., Lanzer, M., Brosius, J., Tang, T. H., & Rozhdestvensky, T. S. (2010). A global view of the nonprotein-coding transcriptome in Plasmodium falciparum. Nucleic Acids Research, 38(2), 608–617. https://doi.org/10.1093/nar/gkp895
Ratti, M., Lampis, A., Ghidini, M., Salati, M., Mirchev, M. B., Valeri, N., & Hahne, J. C. (2020). MicroRNAs (miRNAs) and Long Non-Coding RNAs (lncRNAs) as New Tools for Cancer Therapy: First Steps from Bench to Bedside. Targeted Oncology, 15(3), 261–278. https://doi.org/10.1007/s11523-020-00717-x
Ravishankar, S., Baldelli, V., Angeletti, C., Raffaelli, N., Landini, P., & Rossi, E. (2024). Fluoropyrimidines affect de novo pyrimidine synthesis impairing biofilm formation in Escherichia coli. Biofilm, 7, 100180. https://doi.org/10.1016/j.bioflm.2024.100180
Roth, M., Jaquet, V., Lemeille, S., Bonetti, E.-J., Cambet, Y., François, P., & Krause, K.-H. (2022). Transcriptomic Analysis of E. coli after Exposure to a Sublethal Concentration of Hydrogen Peroxide Revealed a Coordinated Up-Regulation of the Cysteine Biosynthesis Pathway. Antioxidants, 11(4), Article 4. https://doi.org/10.3390/antiox11040655
Downloads
Published
Issue
Section
License
Copyright (c) 2025 Malaysian Journal of Science

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
Transfer of Copyrights
- In the event of publication of the manuscript entitled [INSERT MANUSCRIPT TITLE AND REF NO.] in the Malaysian Journal of Science, I hereby transfer copyrights of the manuscript title, abstract and contents to the Malaysian Journal of Science and the Faculty of Science, University of Malaya (as the publisher) for the full legal term of copyright and any renewals thereof throughout the world in any format, and any media for communication.
Conditions of Publication
- I hereby state that this manuscript to be published is an original work, unpublished in any form prior and I have obtained the necessary permission for the reproduction (or am the owner) of any images, illustrations, tables, charts, figures, maps, photographs and other visual materials of whom the copyrights is owned by a third party.
- This manuscript contains no statements that are contradictory to the relevant local and international laws or that infringes on the rights of others.
- I agree to indemnify the Malaysian Journal of Science and the Faculty of Science, University of Malaya (as the publisher) in the event of any claims that arise in regards to the above conditions and assume full liability on the published manuscript.
Reviewer’s Responsibilities
- Reviewers must treat the manuscripts received for reviewing process as confidential. It must not be shown or discussed with others without the authorization from the editor of MJS.
- Reviewers assigned must not have conflicts of interest with respect to the original work, the authors of the article or the research funding.
- Reviewers should judge or evaluate the manuscripts objective as possible. The feedback from the reviewers should be express clearly with supporting arguments.
- If the assigned reviewer considers themselves not able to complete the review of the manuscript, they must communicate with the editor, so that the manuscript could be sent to another suitable reviewer.
Copyright: Rights of the Author(s)
- Effective 2007, it will become the policy of the Malaysian Journal of Science (published by the Faculty of Science, University of Malaya) to obtain copyrights of all manuscripts published. This is to facilitate:
- Protection against copyright infringement of the manuscript through copyright breaches or piracy.
- Timely handling of reproduction requests from authorized third parties that are addressed directly to the Faculty of Science, University of Malaya.
- As the author, you may publish the fore-mentioned manuscript, whole or any part thereof, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given. You may produce copies of your manuscript, whole or any part thereof, for teaching purposes or to be provided, on individual basis, to fellow researchers.
- You may include the fore-mentioned manuscript, whole or any part thereof, electronically on a secure network at your affiliated institution, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given.
- You may include the fore-mentioned manuscript, whole or any part thereof, on the World Wide Web, provided acknowledgement regarding copyright notice and reference to first publication in the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers) are given.
- In the event that your manuscript, whole or any part thereof, has been requested to be reproduced, for any purpose or in any form approved by the Malaysian Journal of Science and Faculty of Science, University of Malaya (as the publishers), you will be informed. It is requested that any changes to your contact details (especially e-mail addresses) are made known.
Copyright: Role and responsibility of the Author(s)
- In the event of the manuscript to be published in the Malaysian Journal of Science contains materials copyrighted to others prior, it is the responsibility of current author(s) to obtain written permission from the copyright owner or owners.
- This written permission should be submitted with the proof-copy of the manuscript to be published in the Malaysian Journal of Science
Licensing Policy
Malaysian Journal of Science is an open-access journal that follows the Creative Commons Attribution-Non-commercial 4.0 International License (CC BY-NC 4.0)
CC BY – NC 4.0: Under this licence, the reusers to distribute, remix, alter, and build upon the content in any media or format for non-commercial purposes only, as long as proper acknowledgement is given to the authors of the original work. Please take the time to read the whole licence agreement (https://creativecommons.org/licenses/by-nc/4.0/legalcode ).





